Protein Assembly and the Origin of Life
Protein Assembly and the Origin of Life
Abiogenesis is the term used to describe the origin of life from non-life. In modern science, this area is studied by researchers interested in establishing a pathway from raw biochemicals to living cells naturally—without divine assistance. This requires detailed examination of the components of life and complex explanations for how these structures could develop naturally. Unfortunately for those committed to a naturalistic origin of life—one without a Designer—there are huge chemical hurdles that must be overcome–each one seemingly more complex than the previous one.
In their book, The Stairway of Life, Drs. Change Tan and Rob Stadler outline a series of complex steps that must be taken for life to develop.1 They present these steps as a staircase of increasing complexity. Among the questions that must be addressed are how biomolecules originated. The RNA World hypothesis posits that RNA was the first biomolecule, and eventually the ability to build proteins developed from these molecules, with DNA coming along later.
In a recent study, researchers propose a possible solution for a key step toward being able to build proteins in a prebiotic system.2 Their focus was on the step by which amino acids get prepared for being added to proteins. First, let’s review what happens in cells and then explore the solution that has been offered.
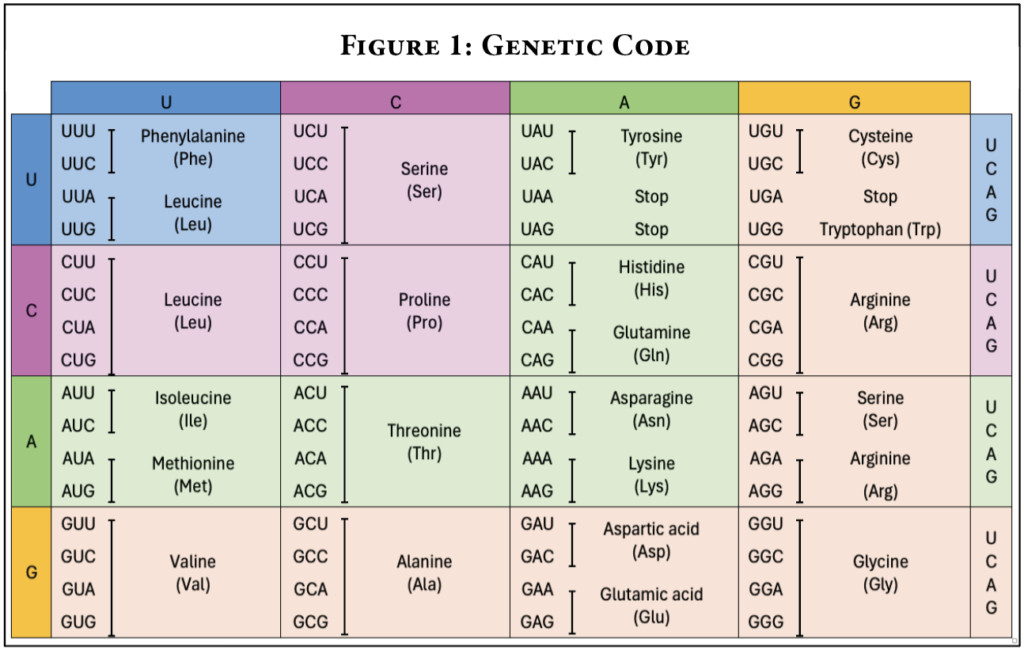
In living organisms, DNA is a storage form for information, and DNA is transcribed into RNA. Here, the term “transcribed” means that the letters of the DNA language are used to arrange the letters of the RNA molecule that is built. Some RNA provides instructions to build proteins. We call this messenger RNA or mRNA. mRNA molecules are used by ribosomes to build proteins in a process called translation. Effectively, the ribosome is translating the nucleic acid “language” in the mRNA into the amino acid “language” of proteins. Sets of three nucleotides in RNA are used to code for one amino acid. The relationship between nucleotides and amino acids is what we refer to as the genetic code (See Figure 1).
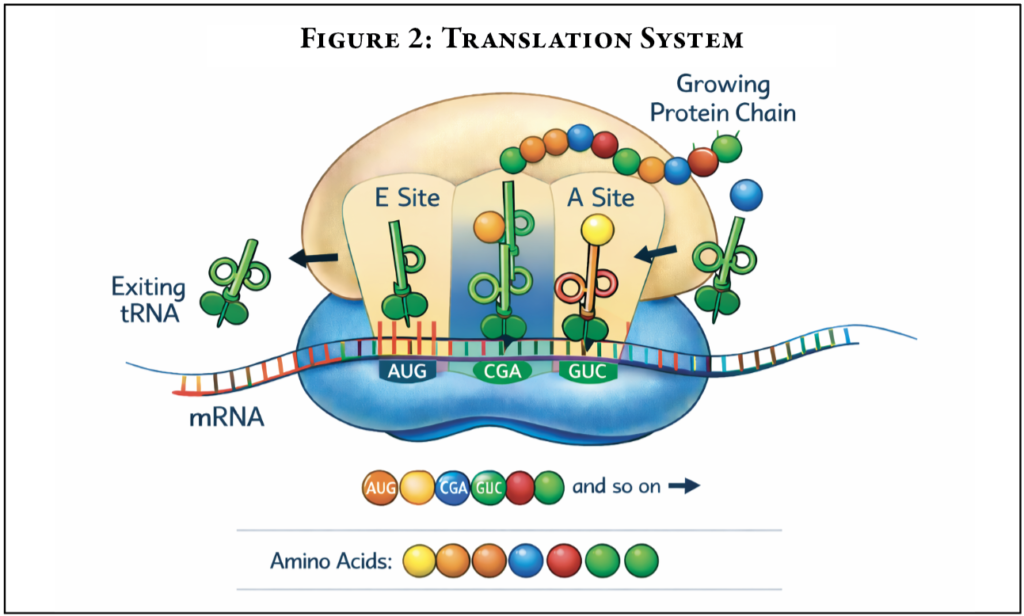
The process of translation is far more complex than what has been described. One major step is the preparation of amino acids to be added to the growing protein chain. In cells, this involves two key players: transfer RNA and aminoacyl-tRNA synthetases. Transfer RNA or tRNA are RNA molecules that deliver amino acids to the ribosome in an “activated” form so the ribosome can build the protein. However, tRNAs are specific for each amino acid–requiring multiple tRNA since there are 20 different amino acids. This requires the correct amino acid to be paired with its corresponding tRNA. This process is carried out by enzymes called aminoacyl-tRNA synthetases (aaRSs). This family of enzymes has a different version for each amino acid. The amino acids and their corresponding tRNAs come together with these enzymes to “charge” the tRNA with the amino acid. The charged tRNA then can deliver the amino acid to the ribosome as it is needed (Figure 2).
In this current study, the researchers realized that you cannot make proteins if you don’t already have aaRSs. So, they propose there are other chemical groups that could have “activated” the amino acids and prepared them for linking in a protein chain. They take advantage of thiol groups or those that contain a sulfur atom as a way to grab and activate an amino acid. They propose a mechanism where this is a pre-step before the aaRS enzymes were available.
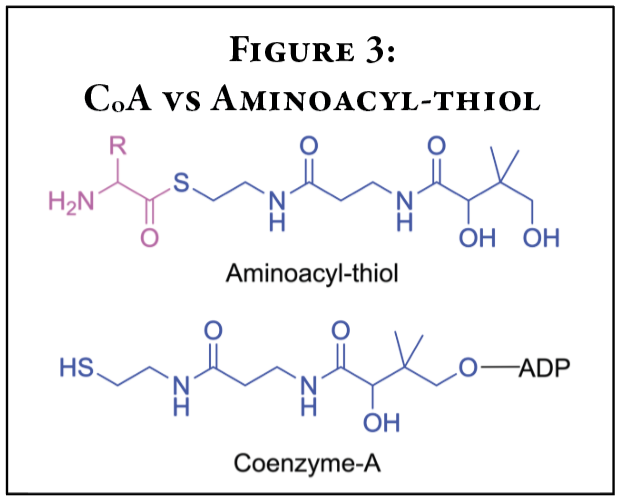
While this is creative chemistry and the thiol groups do exist in cells and in nature, such as in coenzyme-A, this chemistry falls short of explaining how this crucial step could have occurred with specificity for several reasons (Figure 3). First of all, thiol groups are not used in this way currently in our cells. So, there would need to be another explanation for the transition to aaRSs. Second, this chemistry ignores the problem of specificity. While peptides could potentially be assembled, this mechanism provides no explanation for how amino acids could be arranged in specific sequence order. Like the order of letters in this article, the amino acid “letters” must be placed in very specific orders to get functional proteins.
Third, this chemistry utilizes purified, lab-grade compounds under highly controlled conditions. Where would such purified amino acids be available in nature with any abundance prior to the formation of life? How could such a system be sustained over untold thousands to millions of years while “life developed”? Fourth, there is no explanation here for how side reactions could be controlled or suppressed. There are many reactions that had to be closely controlled for these reactions to work. What would control such conditions in natural environments?
Fifth, there is the problem that this chemistry only forms short sequences or peptides–dipeptides and tripeptides. Meaning there are 2 or 3 amino acids together in a chain. This is much smaller than typical proteins, which are often in the range of 300+ amino acids (Figure 4).

Sixth, there is a problem related to the fact that this does not explain the origin of the aaRSs, the tRNA, or the ribosome. Each of these components is essential for living systems at all levels, from bacteria and archaea to all eukaryotes from fungi to plants to animals and humans. So, again, this creative chemistry is interesting, but it does not provide a pathway for the formation of the more complex systems used in living organisms today.
Addressing a larger problem: this research does offer a way to prepare amino acids for being connected together, but it does not give the specificity needed to make real proteins. The system in use in the living world today is highly complex and interdependent. The term used to describe this at times is that it displays causal circularity. As described above, to build a protein, you need amino acids attached to tRNA. To get those, you need proteins referred to as aaRSs to attach each amino acid to its tRNA. So, how do you get aaRS enzymes if you don’t already have them in place since they are proteins? In other words, you need proteins to make proteins. This is true of the ribosome itself. It is made of ribosomal RNA (rRNA) and proteins. How did the first ribosomal proteins get made without a ribosome?
In other words: Which came first, the chicken or the egg? That’s exactly the problem here. To make proteins, you have to have proteins in place already. And not just a few! Protein synthesis requires dozens of proteins, and many of these proteins are complex. Let’s take the aaRSs as an example. These are not small proteins. They typically have 400-1,000 amino acids all in very specific order. How and when did the first aaRSs arrive at such highly specific sequences to carry out their complex function? Again, we need one aaRS for each amino acid–so, we need 20 of these enzymes! That’s just to get the right amino acid ready to go. We have not discussed the large number of proteins involved in transcription (making RNA) and the ribosome. Translation likely requires 80-100 or more proteins to carry out all of the tasks. In addition, the complex pathways needed to build the amino acids require multiple proteins in modern organisms.
In summary, recent research into abiogenesis has offered a possible mechanism for activating amino acids without aaRS enzymes and tRNAs. This mechanism uses a biologically relevant thiol compound and some interesting chemistry to activate amino acids to prepare them for forming peptide bonds. However, it produces short di- and tripeptides that lack sequence specificity, which is required for building large proteins. Thus, this chemistry “trick” does not solve the problems of protein formation in a prebiotic environment. Instead, the research underscores the true complexity of living systems and the need for top-down design of integrated biochemical systems.
Endnotes
1 C.L. Tan and R. Stadler (2020), The Stairway to Life: An Origin-of-Life Reality Check (Sacramento, CA: Evorevo Books).
2 Jyoti Singh, Benjamin Thoma, Daniel Whitaker, Max Satterly Webley, Yuan Yao, and Matthew W. Powner (2025), “Thioester-Mediated RNA Aminoacylation and Peptidyl-RNA Synthesis in Water,” Nature, 644[8078]:933-944, August.

REPRODUCTION & DISCLAIMERS: We are happy to grant permission for this article to be reproduced in part or in its entirety, as long as our stipulations are observed.



0 Comments:
Post a Comment
<< Home